I found out yesterday that Pat Kennedy passed away.
When I met Pat in the early 2000s, he was a quiet, spry gentleman about a decade older than I am now. He was the owner/founder of OSIsoft, the then-largest private software company in the SF Bay Area that you never heard of... unless you work in manufacturing.
Even if you've neither heard of him nor his company, his life's work has probably impacted you. He wrote the software that captures, archives, and retrieves time-series data... instrument data.
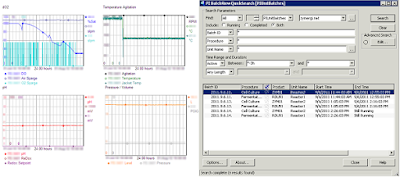
The app opened on nearly every monitor (and the massive display) is PI ProcessBook, the face of the software that he created: PI. And in my world, PI is used everywhere.
At my second job, I was the fermentation engineer at Genentech's Vacaville site. And my first day on the job, my boss - Jesse Bergevin - pulls up PI ProcessBook and tells me that my job is read these squiggles on the screen and divine what is happening in each of the bioreactors.
In 2003, I met Pat for the first time at his annual PI User's Conference in Monterey, California. He rents out the Monterey Bay Aquarium in the evening and throws a huge bash. A group of us were walking back to the hotel all tipsy and I recall asking him why he doesn't take OSIsoft public. He responded simply, "Well, you see, Oliver, if I go public, that means I get a new boss... and to be frank, I don't want a new boss; I like my current boss just fine." This was the first of many words that would shape my worldview.
Our paths would cross again the following year when I was a speaker at one of the conference sessions and of all the talks to attend during my time slot, Pat came to mine!
So I already knew from Herb Boyer & Bob Swanson - the founders of Genentech, that to change the world and get rich (or die trying), you had to be a founder. J. Patrick Kennedy confirmed this for me.
Around the same time, I was reading the book Rich Dad, Poor Dad. In the book, Robert Kiyosaki tells the story of his mentor: Rich Dad, and his father: Poor Dad. That if you live in an environment/mindset you don't like, and you see an environment or mindset that you do like, you need to go find someone with that better mindset and get them to be your mentor. Expecting to get rejected, I fired off a doomed email asking Pat to be my mentor:
From: Oliver Yu
Date: Fri, Jun 24, 2005 at 1:42 PM
Subject: mentor
To: <[email protected]>
Hi Pat -
I am wondering if you would be my mentor. I have been reflecting on where I want to be down the road, and I think that there is no better way of getting to my final destination than to seek the instruction of those who have arrived.
As a start, I'd like to hear you tell your story. I want to know the tough decisions you faced as you founded OSIsoft and the actions you took. How much was instinct, how much was luck? What would you differently if any? And I suppose as I move forward with my pursuits, I'd like to check in with you on your opinion of how I am doing.
I guess for now, I'm asking to buy you dinner and an opportunity to sell you on being my mentor. And if things seem right, we can proceed on a basis that meets your schedule.
Sincerely,
Oliver
Pat responded:
From: Pat Kennedy <[email protected]>
Date: Sat, Jun 25, 2005 at 9:54 AM
Subject: RE: mentor
To: Oliver Yu
Interesting request - can you tell me more about what you want to
accomplish? I am always up for a dinner with a customer, particularly ones that give presentations about how great we are.
Pat
And so, on June 29, 2005, I drove to San Leandro where he gave me words that I couldn't possibly understand. Only decades later did any of that mean anything to me... the few things that stand out in retrospect:
- He never set out to be a founder/software mogul. He was a real estate investor. OSIsoft was his job; real estate was his business.
- He understands the root of his success.
- DuPont had published and given away data compression algorithms.
- His business partner encouraged him to start a consulting firm using these algos.
- He knew to pivot from software to consulting when his customers kept demanding his software and less his consulting
- He understood that his success was a combination of skills, money, luck and serendipity: and that he isn't sure - with immense success - that he could repeat it.
- Picture of Arnold Schwarzenneger standing in the mission control room of the California ISO with PI ProcessBook opened in the background.
- Satellite image of land that he acquired